Camponotus vicinus (Carpenter Ant)
Visual overview of genome assembly metrics
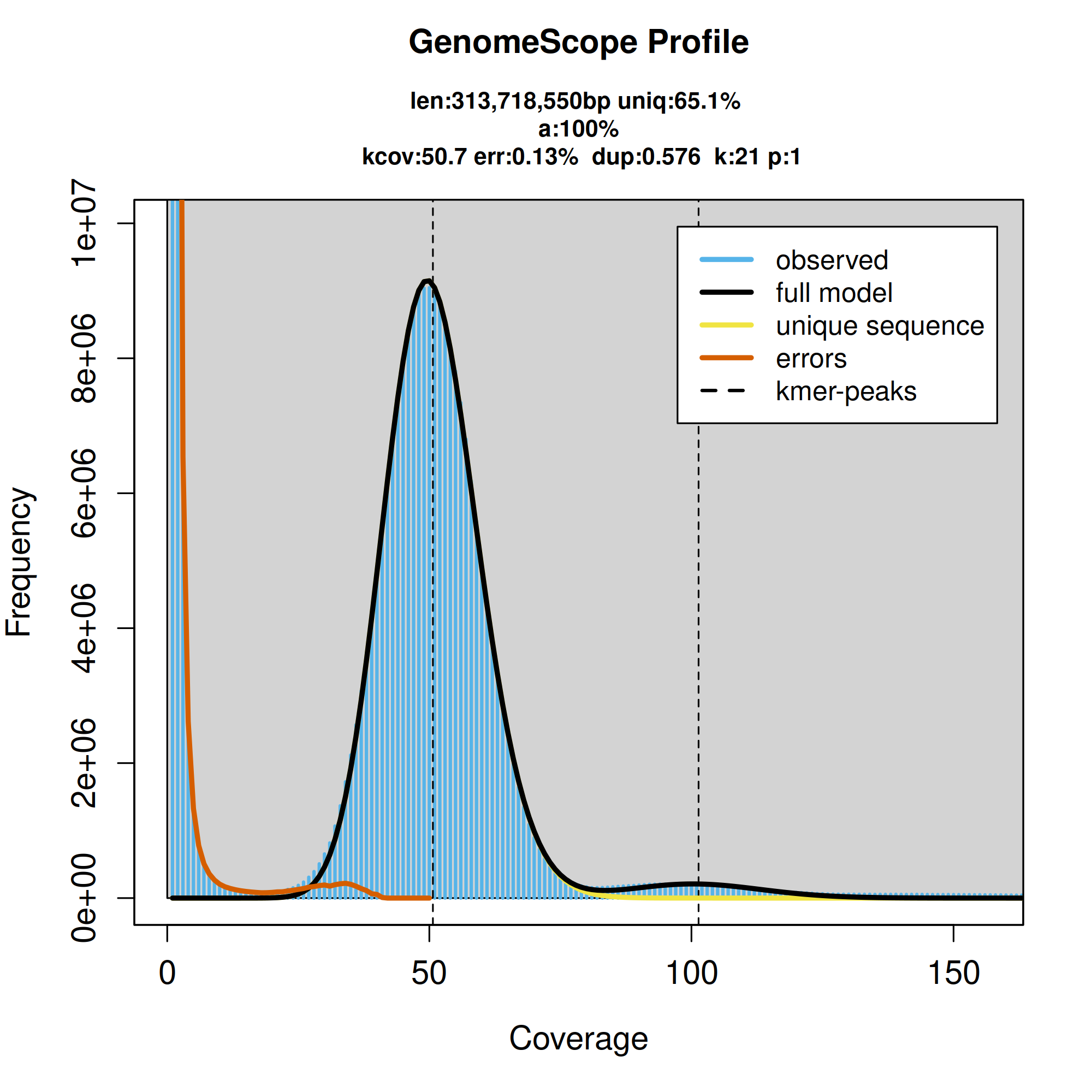
K-mer spectra output generated from PacBio HiFi data without adapters using GenomeScope2.0.
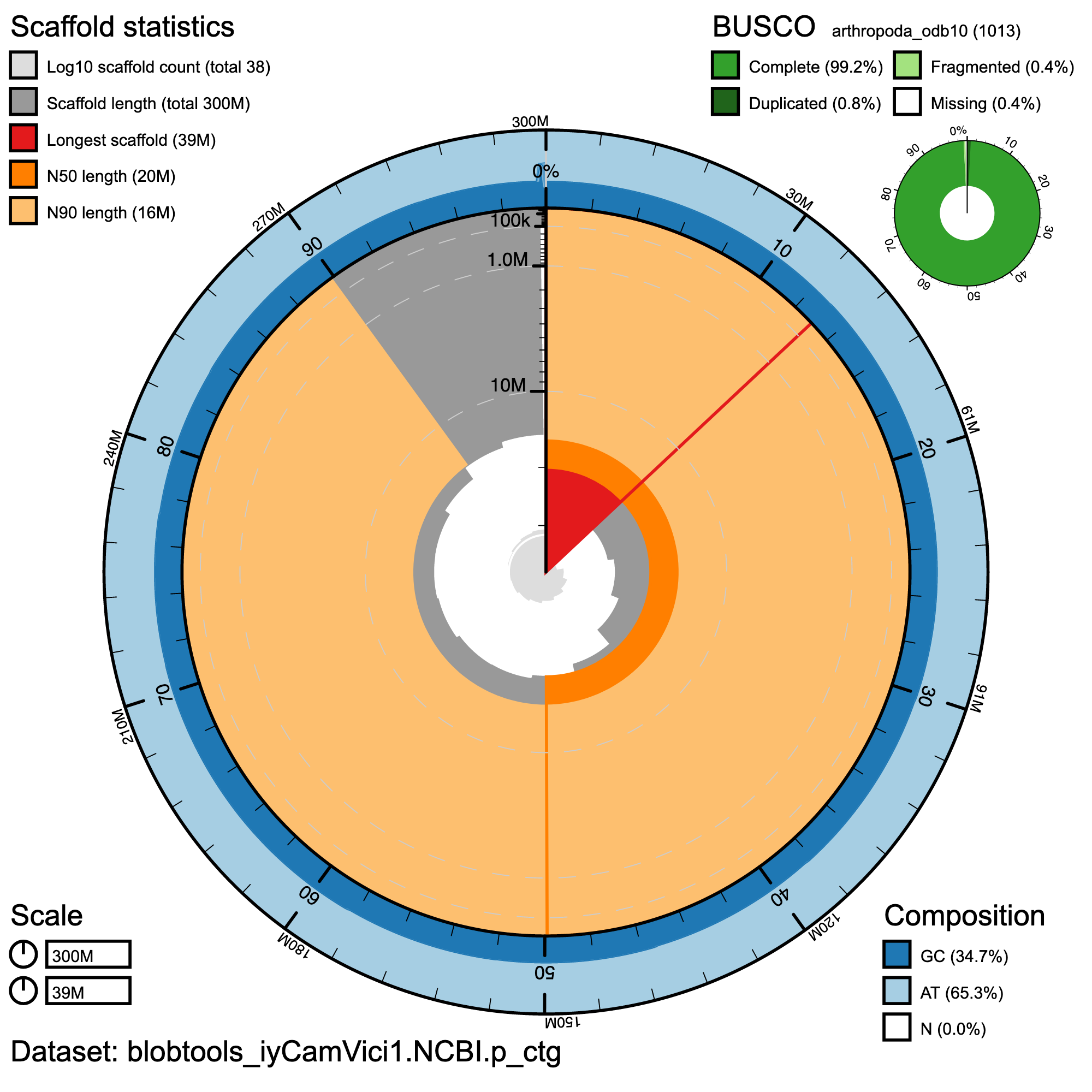
BlobToolKit Snail plot showing a graphical representation of the quality metrics for the primary assembly. The plot circle represents the full size of the assembly. From the inside-out, the central plot covers length-related metrics. The red line represents the size of the longest scaffold; all other scaffolds are arranged in size-order moving clockwise around the plot and drawn in grey starting from the outside of the central plot. Dark and light orange arcs show the scaffold N50 and scaffold N90 values. The central light grey spiral shows the cumulative scaffold count with a white line at each order of magnitude. White regions in this area reflect the proportion of Ns in the assembly The dark vs. light blue area around it shows mean, maximum and minimum GC vs. AT content at 0.1% intervals (Challis et al. 2020)
Omni-C Contact map for the genome assembly generated with PretextSnapshot. The Omni-C contact map translates proximity of genomic regions in 3-D space to contiguous linear organization. Each cell in the contact map corresponds to sequencing data supporting the linkage (or join) between two of such regions. Scaffolds are separated by black lines and higher density corresponds to higher levels of fragmentation.
Authors
Philip S Ward, Elizabeth I Cash, Kailey Ferger, Merly Escalona, Ruta Sahasrabudhe, Courtney Miller, Erin Toffelmier, Colin Fairbairn, William Seligmann, H Bradley Shaffer, Neil D Tsutsui